Creating Source-Localized Time Series from Scalp EEG
function focus on eeg_htpCalcSource.m
Introduction
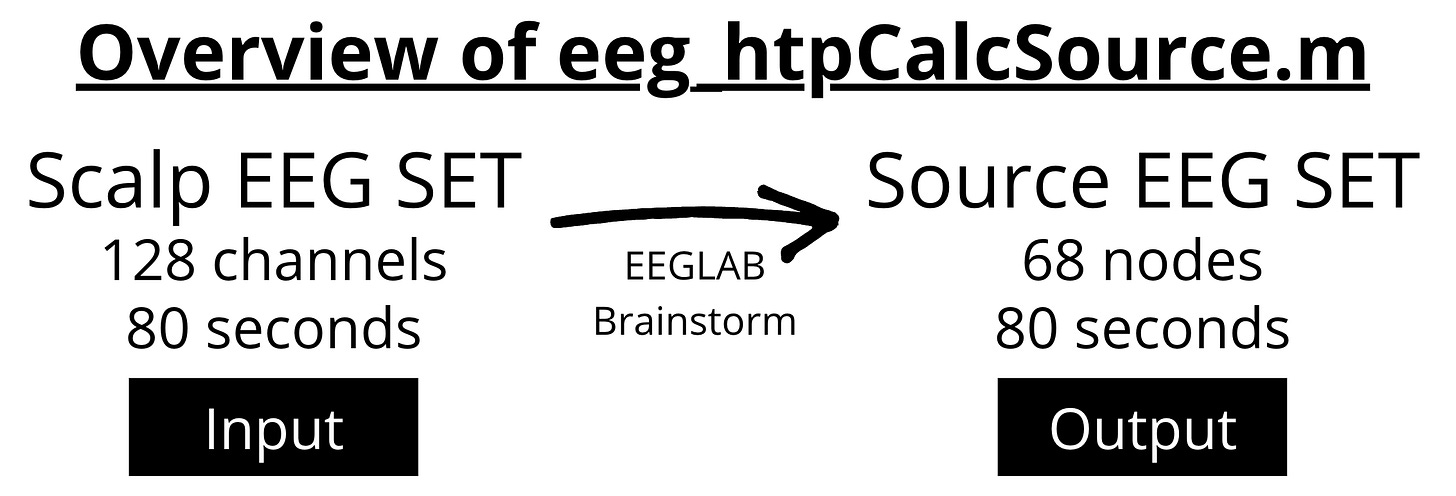
This tutorial covers the basics of using eeg_htpCalcSource.m from the vHTP toolkit for source localization.
Updates:
As of 6/14/2022, the code has been updated:
No longer have to manually specify a headmodel. If you have supported headmodel (EGI128, EGI32, Brainvision64) the correct default headmodel will be downloaded from figshare and automatically imported into the Brainstorm protocol directory. The ‘headmodelfile’ parameter will override any existing headmodel with the file you specify.
Fixed silent crashes by adding an error message if input EEG and headmodel have different number of electrodes.
Verified compatibility with June 2022 release of Brainstorm.
Added demo/testing function with test_eeg_htpCalcSource in the tests directory.
The problem
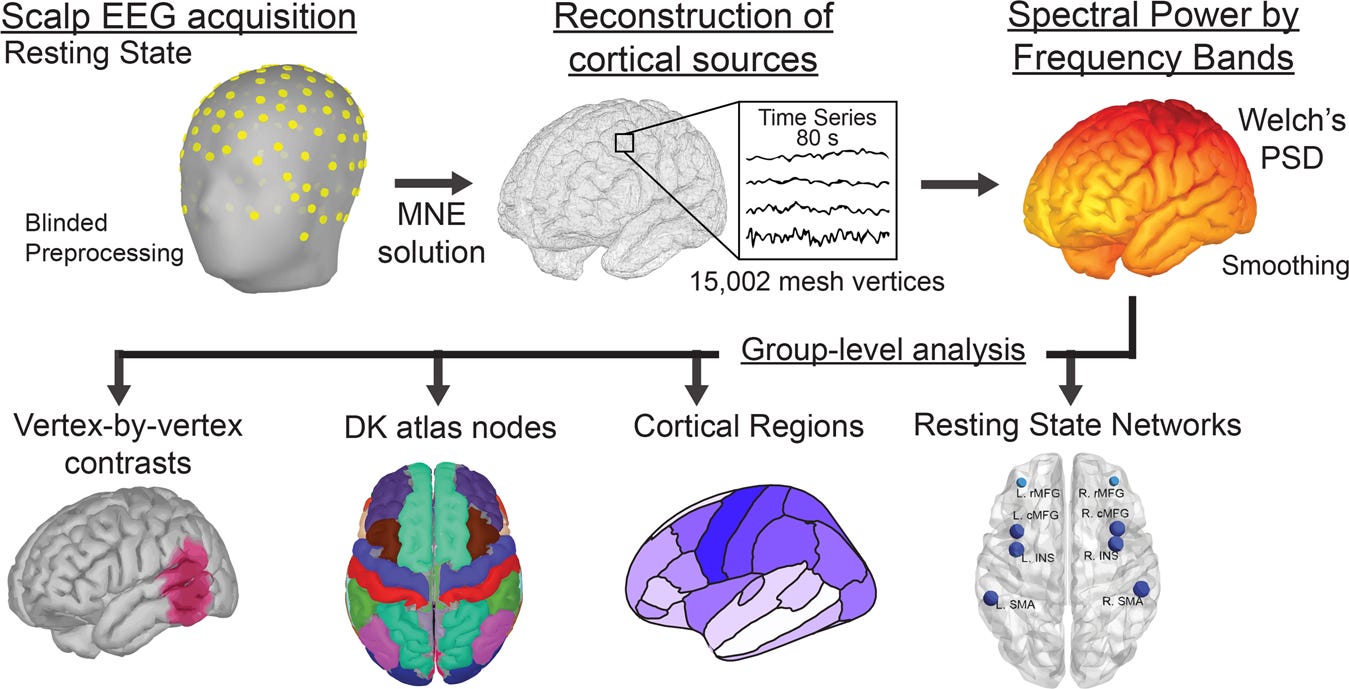
Source estimation resolves EEG sensor scalp amplitude time series into three dimensional ‘source space’ positioned within the brain. There is no perfect method for source localization, nor is one method applicable to every type of analysis. Working out an optimal source solution is not a trivial process, but once you have a workflow how could you reproduce it and consistently apply it to your entire dataset?
A solution
Once configured the eeg_htpCalcSource function can accomplish this in a single step using EEGLAB and Brainstorm command line functions:
The input of the function is an artifact-free EEGLAB SET file of scalp-sensor EEG data. Following the process, the source estimation will also be available in the Brainstorm graphical interface for visualizing accuracy and additional analysis.
Dependencies
Several open source toolkits are used with vHTP.
Download following toolkits:
Each toolkit should be installed within a separate directory (per software instructions) and added to the MATLAB path.
EEGLAB: https://github.com/sccn/eeglab
Brainstorm: https://github.com/brainstorm-tools/brainstorm3
Run each toolkit for initial setup
Run EEGLAB from the MATLAB command prompt using
eeglabRun Brainstorm from the MATLAB command prompt using
brainstorm
You will need sufficient room to generate source files. Select data directories that can handle several gigabytes of EEG data (including duplicate files). Brainstorm requires a separate data directory:
Tutorial Parameters
For each set of scalp data and analysis you will likely need to consider parameters optimal for your situation. For this tutorial, we will use a process that we used for a recently published manuscript using source localization to characterize EEG signals in Fragile X Syndrome:
Pedapati, E.V., Schmitt, L.M., Ethridge, L.E. et al. Neocortical localization and thalamocortical modulation of neuronal hyperexcitability contribute to Fragile X Syndrome. Commun Biol 5, 442 (2022). https://doi.org/10.1038/s42003-022-03395-9
Input data specifications:
For this tutorial, we have sample cleaned resting EEG file which was recorded on an EGI NetAmp 400 with 128-channel Hydrocel saline based geodesic style nets. To learn more about why this particular configuration is well-suited for source localization see:
Song, Jasmine, et al. "EEG source localization: sensor density and head surface coverage." Journal of neuroscience methods 256 (2015): 9-21.
Dataset: The resting state sample file can be downloaded from here. We will have an upcoming blog post detailing using vHTP for preprocessing EEG data.
The preprocessing for this data is described in detail in Pedapati et al., but briefly data was resampled to 500 Hz (original 1000 Hz) filtered between 2 and 90 Hz, notch filter for line noise, bad channels and continuous artifact removed, underwent artifact subspace reconstruction, and ocular/heart rate/and muscular artifacts removed with ICA. No individual MRI or electrode positions are available for this subject.
Downloading a precomputed head model
The projection of electrode sources is accomplished through a head model which can estimate electric fields based on the geometry of the electrode positions and the conductivity of tissue.
When individual MRIs and digitized electrode positions are not available, a default head model created from an averaged structural MRI and standardized electrode positions can be used. It is computationally intensive to generate this model, thus, when using default anatomy a single head model can be generated and copied for use by each subject (of course, each scalp EEG should be identically recorded on the same equipment). A Brainstorm forum post here describes this process.
We have used the OpenMEEG to generate a default head model for the specific electrode net used for our scalp recordings. While working through this tutorial, you will need to download this file and make it available for the script to use.
Depending on your use case, you can create other default head models based on a specific EEG nets or even generate individual models (with slight modification to the code). A simple loop using the ‘headmodelfile’ parameter can specify the specific head model file to use.
Update 6/13/2022: The newest version of the code includes automatic downloading of several common head models.
Computing a new head model
If you are using individual MRI or a different (or captured) channel montage, you may want to generate a new head model.
Specify the parameter ‘computeheadmodel’ as true. This will compute the headmodel and add the file to default directory. The file will then be copied to other subjects. We are working on a version that will allow for individual MRI/electrodes shortly!
Checking the prerequisites
A complete setup will include:
functioning EEGLAB and Brainstorm installations.
The vHTP toolbox should also be available on the path.
The sample dataset should be placed in a data directory.
Finally, the head model file can also be placed in the same folder as the data.
Setting up the script (updated)
% Creating Source-Localized EEG Time Series
% June 8, 2022
% Dependencies - check if toolkits installed
eeglab nogui;
brainstorm;
% setup directories
datadir = '/srv/RAWDATA';
samplefile = 'D0079_rest_postcomp.set';
% Load sample datafile
EEG = pop loadset ('filepath', datadir, 'filename' samplefile);
% Force double precision
EEG.data = double(EEG.data);
% Perform source estimation using Brainstorm
sEEG = eeg_htpCalcSource( EEG );
% Save source time series
pop saveset (sEEG, 'filepath', datadir, 'filename' SEEG.filename)
Let’s walk through this example script:
The initial few lines confirm the presence of required toolboxes.
Next we setup specific path information: data directory and data filename.
We load the dataset and force double precision
Brainstorm prefers a continuous strip of data when working with resting state, this will be performed within the source function.
Next, we run the vHTP function to generate the source model in Brainstorm. The output variable SEEG contains the source time series (68 nodes).
The source localized SET file is saved into the directory for later use. We have opted to use the DK atlas for scout time series.
Locating the source model in Brainstorm
The newly created source file will exist in Brainstorm under the custom protocol eeg_htp. As the data has been formally loaded into Brainstorm, you can perform brainstorm operations on the dataset - including recomputing a source model with different parameters. You can also confirm electrode positions and labels.
New EEG source localized time series
The output the function is an EEG SET structure which conveniently contains timeseries of each atlas ROI instead of electrode channels.
Conclusion
In this blog, we looked at a sample workflow of creating source-localized time-series set files which can be easily implemented into other analysis routines.